Read the most recent reviews and papers: 385, 398, 401, 409, 410
Modern computational analysis has provided chemists the power to rapidly perform complex calculations, leading to increased understanding and expedited optimization of chemical systems. An important application of this technology is its implementation in catalyst design. Currently, intensive quantum mechanical calculations can be routinely performed to generate optimized geometries and energies of reactive intermediates and transition states. However, despite the power of transition state modeling to aid in the design of new catalytic entities, the reliance on chemical intuition to do so is inherently flawed. No matter how experienced or insightful the practitioner, several problems conspire to hinder rapid progress: (1) the lack of detailed mechanistic understanding of the rate and stereodetermining events in a process, (2) the inherent limitations of the human brain to find patterns in large collections of data, and (3) the lack of quantitative measures to aid the choice of catalyst candidates. Moreover, current computational techniques work retrospectively to explain experimental observations. Because of the high computational demands of quantum mechanical calculations, it is not feasible to use these techniques predictively to guide catalyst design. Further, as mentioned above, the use of this approach requires a thorough mechanistic understanding of the transformation in question.
Our chemoinformatics approach provides an attractive alternative because: (1) no mechanistic information is needed as the substrates are not included in the analysis, (2) catalyst structures are characterized by 3D-descriptors that quantify the steric and electronic properties of thousands of candidate molecules, and (3) the suitability of a given candidate can be quantified by comparing its properties to a computationally derived model based on experimental data. This kind of analysis, known as quantitative structure activity relationships (QSAR), is popular in the pharmaceutical industry for understanding the activity therapeutically relevant molecules. The application of QSAR to catalysis is conceptually similar except that the correlation of molecular properties is made to a chemical transformation rather than a binding event or inhibition of a biological process.
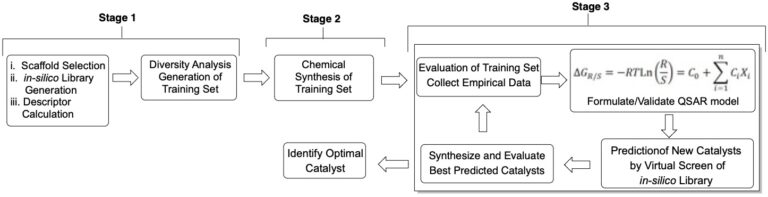
We have recently introduced a fully chemoinformatics-guided workflow to accomplish this goal. The workflow consists of the following components: (1) a large, in silico library of synthetically accessible catalyst candidates is constructed; (2) for each member of this library, conformer dependent descriptors are calculated which define the chemical space of the library; (3) from this library, a representative subset is algorithmically selected, termed the Universal Training Set (UTS) because it is selected considering only catalyst properties and is thus agnostic to reaction and mechanism; (4) this UTS is synthesized and evaluated in the reaction of interest; (5) statistically-validated mathematical models are constructed relating the calculated descriptors to experimental outcome; (6) the in silico library is virtually screened with the model and the best catalyst candidates (along with confidence metrics) for the particular transformation are identified for synthesis; and (7) experimental validation of the prediction. This process can be performed iteratively, with each subsequent iteration added to the training data, until an ideal catalyst is identified.
Full Chemoinformatic Workflow:
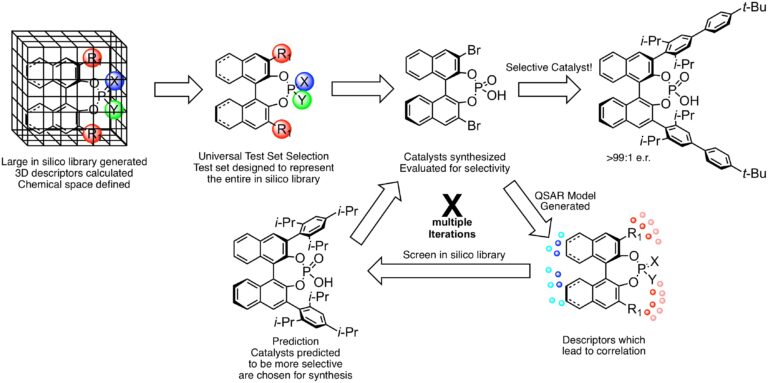
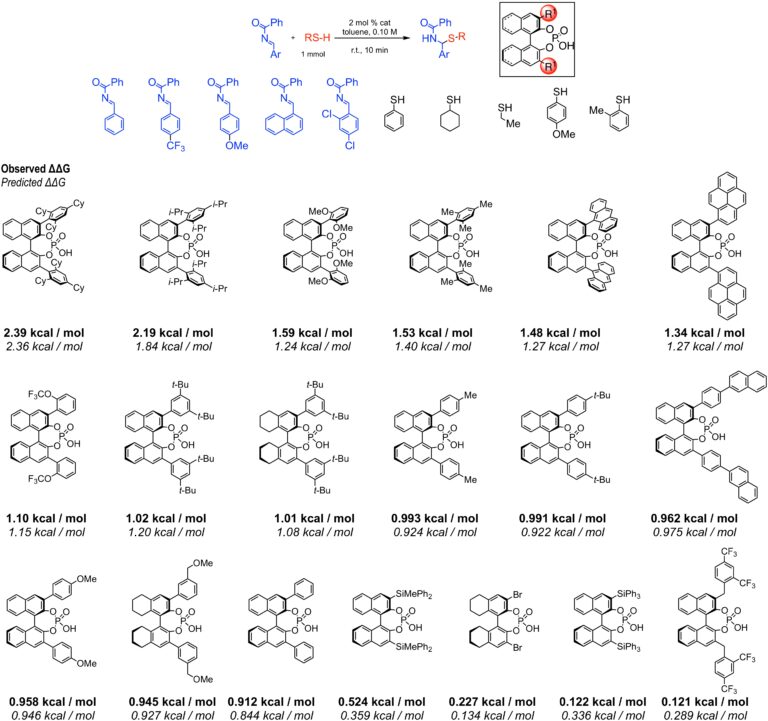
To demonstrate this potential in our method, we: (i) predicted reaction outcomes with substrate combinations and catalysts not used in the training data, and (ii) simulated a situation in which highly selective reactions have not been achieved. The reaction employed to illustrate these predictions was the enantioselective addition of thiols to acyl imines catalyzed by BINOL phosphoric acids. In the first demonstration, a model was constructed using support vector machines and was validated with three different external test sets with MADs ranging from 0.161 to 0.0236 kcal/mol. In the second study, no reactions with selectivity above 80% ee were used as training data. Deep feedforward neural networks accurately reproduced the experimental selectivity data, successfully predicting the most selective reactions. More importantly, the general trends in selectivity, on the basis of average catalyst selectivity, were correctly identified. Despite omitting approximately half of the experimental free energy range from the training data, accurate predictions could still be made in this region of selectivity space.
Observed and Predicted Selectivities of Test Catalysts for All Substrate Combinations:
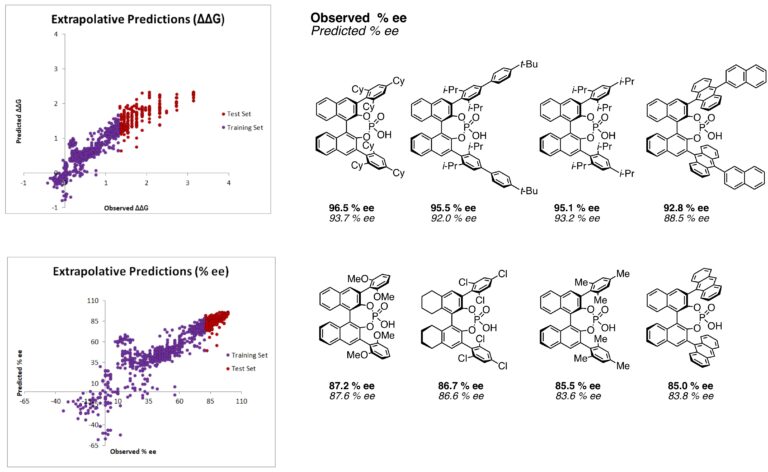
Predicted Selectivities >80% ee Using <80% ee Training Data:
Future implementations of this workflow involve the optimization of other transformations catalyzed by chiral Brønsted acids as well as the development of libraries and training sets for other privileged catalyst scaffolds (bisoxazolines, phosphino-oxazolines, TADDOLs, etc.).
